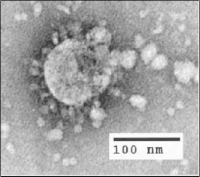
Photo from wikipedia
Significance The ongoing COVID-19 pandemic is an urgent, global threat. Here we analyze the genome of the virus that causes COVID-19, SARS-CoV-2, along with other members of the coronavirus family.… Click to show full abstract
Significance The ongoing COVID-19 pandemic is an urgent, global threat. Here we analyze the genome of the virus that causes COVID-19, SARS-CoV-2, along with other members of the coronavirus family. Our analysis identifies crucial genomic features that are unique to SARS-CoV-2 and two other deadly coronaviruses, SARS-CoV and MERS-CoV. These features correlate with the high fatality rate of these coronaviruses as well as their ability to switch hosts from animals to humans. The identified features could represent crucial elements of coronavirus virulence, and allow for detecting animal coronaviruses that have the potential to make the jump to humans in the future. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) poses an immediate, major threat to public health across the globe. Here we report an in-depth molecular analysis to reconstruct the evolutionary origins of the enhanced pathogenicity of SARS-CoV-2 and other coronaviruses that are severe human pathogens. Using integrated comparative genomics and machine learning techniques, we identify key genomic features that differentiate SARS-CoV-2 and the viruses behind the two previous deadly coronavirus outbreaks, SARS-CoV and Middle East respiratory syndrome coronavirus (MERS-CoV), from less pathogenic coronaviruses. These features include enhancement of the nuclear localization signals in the nucleocapsid protein and distinct inserts in the spike glycoprotein that appear to be associated with high case fatality rate of these coronaviruses as well as the host switch from animals to humans. The identified features could be crucial contributors to coronavirus pathogenicity and possible targets for diagnostics, prognostication, and interventions.
Journal Title: Proceedings of the National Academy of Sciences of the United States of America
Year Published: 2020
Link to full text (if available)
Share on Social Media: Sign Up to like & get
recommendations!